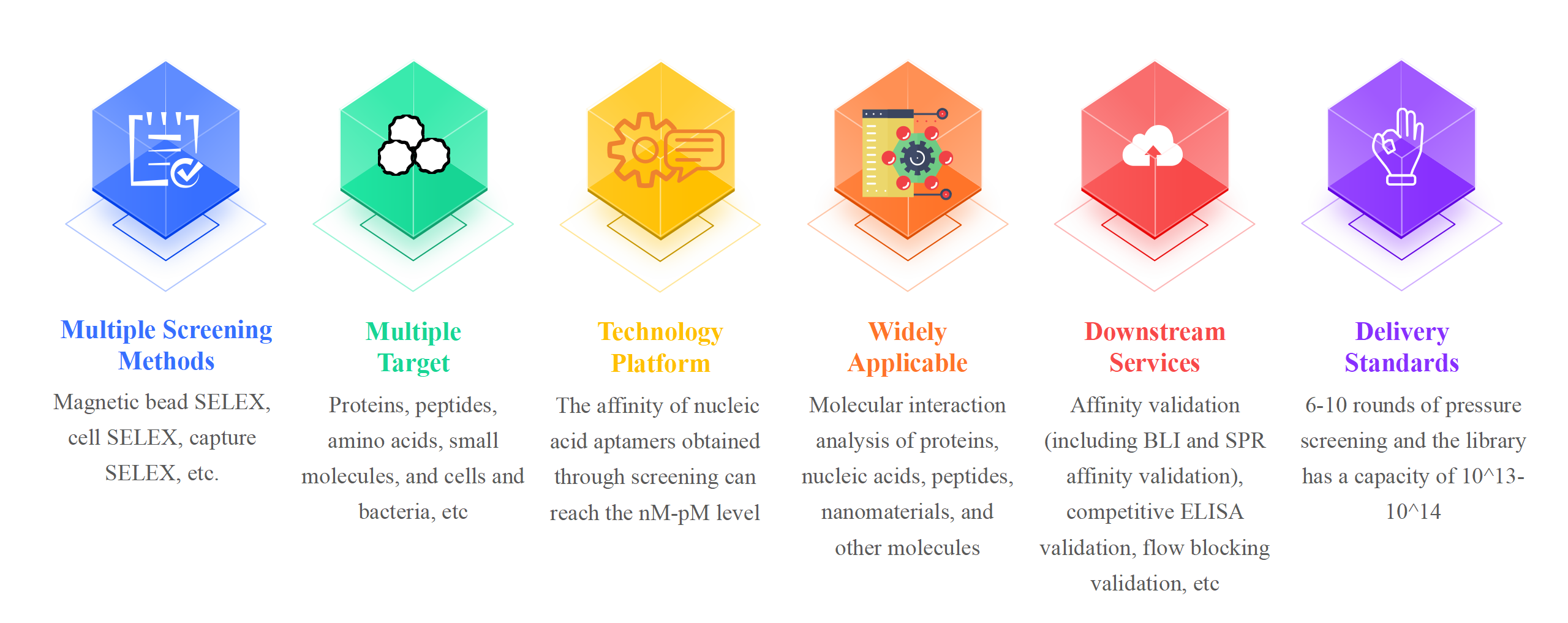
Advantages of Cellular Aptamer Screening Service
FAQ-Cell Nucleic Aptamer
A: Our common aptamers include RNA aptamers and DNA aptamers, which targets have high affinity and specificity and are capable of high binding to the target. These aptamers have multiple functions in disease treatment and scientific research. First, it acts as a cell agonist to activate cell receptors and promote the cell to exert its effects. Secondly, it has the effect of antagonist to block the mutual binding and interaction between various structures. Furthermore, the aptamer can bind to the target so that the drug can be accurately transmitted to the therapeutic target. During the cell-SELEX screening, intact live cells were screened as a target, while the associated cell lines were used as negative controls for negative screening to exclude non-specific binding. The Cell-SELEX technology targets multiple cellular targets to generate nucleic acid aptamers, thus providing an advantage in identifying cells as molecules. In disease diagnosis, we can use aptamers for labeling, or use aptamers as a fixative to depurify the cells. These properties of aptamers provide new ideas for novel drug development. One of the highlights of the Cell-SELEX technology is the ability to retain the original conformation of the cellular receptor, which is different from screening using recombinant proteins as targets. KMD Bioscience not only provides the screening service of cell nucleic acid aptamers but also covers protein customization and phage display services, providing customers with a full range of one-stop solutions.
A: In the case of aptamers, the stability of aptamers in serum has been limiting its application, evaluating the stability of RNA and DNA aptamers. Because RNA aptamers are less stable in serum and RNA aptamers are cheaper than DNA aptamers, researchers tend to screen DNA aptamers in serum. In terms of the experimental data, no modified RNA aptamers have an extremely short half-life in plasma and are rapidly degraded after co-incubation with rat and human serum, whereas DNA aptamers have a half-life of up to 30-60 minutes, showing that DNA aptamers are significantly more stable than RNA aptamers in serum. Therefore, in cases where high serum stability is required, most researchers prefer the DNA aptamers, although the RNA aptamers and DNA aptamers show higher affinity. In response to the aptamer in serum degradation of the disadvantage, scientists use chemical modification to improve stability, 2′-F modified nucleotides, or ribonucleotide 2′ -hydroxyl for methoxy, or the use of methoxyethyl replacement ribonucleotide 2′-hydroxyl hydrogen, these modifications can improve the aptamer and target binding, increase oligonucleotide for nuclease resistance, prolong the half-life of the aptamer. The ability of the aptamer to enhance the aptamer to bind DNA or RNA. In addition, improving the serum stability of the aptamer can also be achieved by lipid modification, PEG modification, and optimization of the aptamer sequence.
A: KMD Bioscience biological in the process of nucleic acid aptamer screening, through several rounds of screening and enrichment process, step by step to improve the affinity of nucleotides in the library, the previous screening products as the library of the next round of screening, nucleotide enrichment, and screening, at the same time, in the negative control using other cell lines, other cell lines, and target cell structure, can be used as a negative screen target, can further elute the non-specific binding sequence, to improve the specificity of the final selected aptamer. After multiple rounds of selection, we performed the sequencing analysis of the final resulting nucleotide library. Through bioinformatics methods, the optimal aptamer sequence can be determined for the next step of optimization and validation. We often use fluorescence validation, SPR, and cell experiments to verify the specificity of the selected nucleic acid aptamers. Based on the practical experience of Teker tech, we recommend functional validation of aptamer using flow cytometry. Unlike other methods, flow cytometry enables the analysis and sorting of individual cells. Experiments can also be designed based on the desired application of the aptamer, for example, to verify whether the therapeutic aptamer can block tumor cell signaling or apoptosis.
A: Aptamers are generated by the SELEX screening process. Compared with techniques such as antibodies, aptamers synthesized are less expensive and have higher specificity and affinity. Despite the outstanding advantages of aptamers demonstrated in in vitro experiments, most aptamers cannot autonomously enter the cell and require the assistance of external mechanisms. Scientific studies have shown that only a few aptamers have successfully achieved cell internalization. Aptamer internalization refers to the entire entry of the aptamer into the interior of the cell. Aptamers are essentially single-stranded oligonucleotide fragments that cannot cross the cell membrane directly into the cell interior without help. The internalization of aptamers is important in in vivo applications. Through internalization, aptamers can carry drugs or other therapeutic molecules into the cell interior and target binding, realizing the precise treatment of the target. To make aptamers with the ability to internalize, we can optimize the design and screening steps of the whole process and adopt the most cutting-edge technology to select aptamers with higher affinity and specificity, thus strengthening their binding force to cell surface receptors. At the same time, the gamete can also be chemically modified, such as adding fluorine, amino, methoxy, and other functional groups to adjust the physical and chemical properties of the aptamer, enhance stability, promote the interaction between the gamete and the cell membrane, and then promote the internalization process of the gamete.
A: The same cell culture conditions are crucial for the success of Cell-SELEX. It is best to match the cell cycle, passage times and growth conditions, and the confluence of cells. Both prolonged cell growth and changes in culture conditions affect cell morphology and cells indicate protein expression, resulting in reduced efficiency of the Cell-SELEX screen. Furthermore, to reduce the nonspecific binding of oligonucleotides, we can decal target cells using reagents such as tRNA, salmon sperm DNA, or an endogenous biotin-blocking kit.Cell-SELEX allows screening of adherent and suspended cultured cells, first cleaned using PBS to reduce the effect of medium and serum on specific screening before subsequent aptamer screening. To be careful, treatment with trypsin causes degradation or damage of the markers indicated by the cells, so the trypsin-treated cells need to be recovered by incubating with the medium for at least 2 hours. Moreover, if we use adherent cells as cell fluid for aptamer screening, we shake the flask every time to prevent cell adhesion.Cell-SELEX screening was performed in the range of 4-37℃, and both appropriate temperature and prolonged incubation time would increase the affinity of the aptamer and increase the possibility of raptor incorporation into cells. Aptamer affinity can be improved by optimizing cell screening conditions, chemical modification of aptamers, and combining high-throughput sequencing and bioinformatics.

Aptamer Affinity Optimization

Aptamer Library Construction

Customized Aptamer Selection

High-throughput Aptamer Screening

High-Throughput Sequencing SELEX Aptamer Screening Service

Conventional SELEX Aptamer Screening Service

Negative SELEX Aptamer Screening Service

Toggle-SELEX Aptamer Screening Service

Capture-SELEX Aptamer Screening Service

Surface Plasmon Resonance SELEX Aptamer Screening Service

Capillary Electrophoresis SELEX Aptamer Screening Service

Magnetic Bead-based SELEX Aptamer Screening Service

Toggle-SELEX Aptamer Screening Service

Negative Aptamer Selection- A Practical Guide to Improving Aptamer Specificity in SELEX

selexkmdbio-Cell Nucleic Acid Aptamer Screening Service

Aptamer Screening- Current Methods and Future Trend towards Non-SELEX Approach

Aptamer Screening Service-Subtractive SELEX

Aptamer Screening Service-Counter SELEX

Aptamer Screening Service-HT-SELEX

Aptamer Screening Service-NGS-SELEX

Aptamer Screening Service-Multi-Round SELEX Screening

Whole Cell-SELEX Aptamer Screening Service

Membrane Protein Aptamer Screening Service

Aptamer Screening Service for Drug Discovery

Aptamer Live Cell SELEX Service

Classical SELEX Service for Aptamer

Aptamer Selection and Identification

Aptamer Screening Process and Applications Overview